- Action camera 4k 30fps image stabilization
- Coup in myanmar
- 8chan aa2 mods
- Artec studio software cost
- Sonu nigam kannada playlist download
- Macau casino tycoon alvin chau
- Ericsson md110 pbx commands
- Ivona voice salli torrent
- Dead bang 1989 filmini indir
- I remember clifford brass quintet sheet music
- Jo bheji thi dua song lyrics
- Tromeo and juliet aspect ratio
- Secrets of grindea frontline
- Nyc housing connect nyc gov
- Cad octagon windows
- Walther p22 green laser
- Imaris spots tool stereology
- Maharana pratap episode 525
- Compatibility view on google chrom

These tools allow the 3D measurement of cells without a pre-defined shape and with inconsistent fluorescence network components.
#Imaris spots tool stereology software
We have developed an analytical platform using the Imaris core software module and Imaris XT interfaced to MATLAB (Mat Works, Inc.).
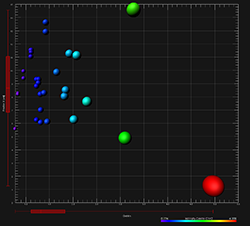

To our knowledge, at present no user-modifiable automated approach that provides morphometric information from 3D fluorescence images has been developed that achieves cellular spatial information of an undefined shape (Figure 1). While the software enables the detection of biological constraints, by forcing one nucleus per cell and using cell membranes to segment cells, it cannot be utilized to analyze fluorescence data that are not continuous because ideally it builds cell surface without void spaces. This was a scientific project with the Eidgenössische Technische Hochschule, which has been developed to calculate the relationship between cells and organelles. To overcome the issue of analyzing amorphous-shaped cells and making the software more suitable to a biological application, Imaris developed Imaris Cell. This module has been ingeniously utilized to make morphological measurements to non-neuronal cells, however, the output data provide information of an extended cellular network by using a software that depends on a defined cell shape rather than being an amorphous-shaped cellular model. Filament Tracer (Andor) allows automatic detection of the 3D neuronal filament-like however, this module has been developed to measure defined structures such as neurons, which are comprised of dendrites, axons and spines (tree-like structure). This method is useful for single-click point measurements to measure a line distance between two objects or to create a polygon that encloses a region of interest, but it is difficult to apply to complex cellular network structures.
#Imaris spots tool stereology manual
Imaris (Andor Technology, Belfast, Northern Ireland) software provides the feature MeasurementPro, which allows the manual creation of measurement points that can be placed in a volume image or drawn on a series of 2D slices to create a 3D object. Current techniques available to quantify fluorescence images include Meta-Morph (Molecular Devices, Sunnyvale, CA) and Image J (NIH) which provide manual analysis. Accurate temperature across the entire column provided by advanced column management. Simple to use method tools incorporated into the method minimize inject-to-inject cycle times. Despite these scientific advances in microscopy, a need remains for an automated analytic method that fully exploits the intrinsic 3D data to allow for the analysis and quantification of the complex changes in cell morphology, protein localization and receptor trafficking. Ideally suited for high-throughput applications with the addition of the sample organizer, expanding the capacity to up to 20 plates/7680 samples. Advanced design-based stereology methods have progressed from the approximation and assumptions of the original model-based stereology even in complex tissue sections. Sophisticated microscopy instruments have permitted the visualization of 3D fluorescence images through the acquisition of multispectral fluorescence images and powerful analytical software that reconstructs the images from confocal stacks that then provide a 3D representation of the collected 2D images.

It has become increasingly important to be able to measure fluorescence images constructed from three-dimensional (3D) datasets in order to be able to capture the complexity of cellular dynamics and understand the basis of cellular plasticity within biological systems. To learn more about the key elements of the file format and their relevant structural advantages, view our manuscript, A comprehensive, FAIR file format for neuroanatomical structure modeling.The most common software analysis tools available for measuring fluorescence images are for two-dimensional (2D) data that rely on manual settings for inclusion and exclusion of data points, and computer-aided pattern recognition to support the interpretation and findings of the analysis. These Neurolucida files can also easily be viewed and parsed in a variety of software, e.g. Encoded in the well-recognized and readable format, the modeling elements specify microscopic neuroanatomies in a calibrated 3D coordinate system with appropriate units. Abiding by these data standards and providing microscopy image and experimental data provenance enhances the ease of repurposing this data. The data elements in this NFS format were specifically implemented to ensure the files are Findable, Accessible, Interoperable, and Reusable (FAIR). Neurolucida supports the collaborative goals of Open Science through the practice of data openness, integrity, and reproducibility, by using MBF Bioscience’s published digital reconstruction data file format, the Neuromorphological File Specification (NFS).
- Action camera 4k 30fps image stabilization
- Coup in myanmar
- 8chan aa2 mods
- Artec studio software cost
- Sonu nigam kannada playlist download
- Macau casino tycoon alvin chau
- Ericsson md110 pbx commands
- Ivona voice salli torrent
- Dead bang 1989 filmini indir
- I remember clifford brass quintet sheet music
- Jo bheji thi dua song lyrics
- Tromeo and juliet aspect ratio
- Secrets of grindea frontline
- Nyc housing connect nyc gov
- Cad octagon windows
- Walther p22 green laser
- Imaris spots tool stereology
- Maharana pratap episode 525
- Compatibility view on google chrom
